August 8, 2022 — Representatives from the Diagnostics Working Group of the IASLC Early Detection and Screening Committee ...
Clinical Trials
Clinical trials and studies about imaging technology can be found on this channel.
August 8, 2022 — A clinical trial is only as powerful as its participants. For years, researchers have struggled to fill ...
July 27, 2022 — Multitasking is not just an office skill. It’s key to functioning as a human, and it involves something ...
July 25, 2022 — For any drug to be successful in treating mental health or neurological problems, the medicine has to ...
July 20, 2022 — Whiterabbit.ai, an AI company targeting late-stage diseases, today announced efforts to recruit ...
July 19, 2022 — Hebrew University of Jerusalem (HU) researchers have developed an early-stage diagnosis of Parkinson’s ...
July 13, 2022 — KA Imaging, a company that develops innovative X-ray imaging solutions, announced the initial results ...
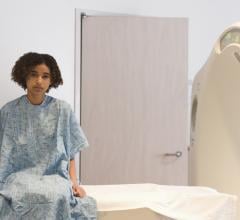
July 6, 2022 — Lung cancer is the most common cancer in the world, with 2.2 million new cases and around 1.8 million ...
June 29, 2022 — Building on Penn Medicine’s years of research and use of imaging technology that illuminates tumor ...
June 27, 2022 — Varian, a Siemens Healthineers company, announced that the U.S. Food and Drug Administration (FDA) ...
June 27, 2022 — Neutron Therapeutics, Inc (NT) and the University Hospital of Brussels (H.U.B) today announced that they ...
June 24, 2022 — Recently, a collaborated research team led by Prof. LI Hai and Hongzhi Wang from Hefei Institutes of ...
June 7, 2022 — Two-year results from the Dysphagia-Aspiration Related Structures (DARS) trial, which is funded by Cancer ...
June 1, 2022 — According to ARRS’ American Journal of Roentgenology (AJR), evaluating chest CT findings offers an ...
May 31, 2022 — Patient trust is not a barrier to artificial intelligence (AI) adoption by medical imaging professionals ...
Clinical scientists used machine learning (ML) models to explore de-identified electronic health record (EHR) data in ...
May 13, 2022 — Alzheimer’s disease is the most common form of dementia, one of the major causes of dependency and ...
May 12, 2022 — Abbott announced a new partnership with Women as One to launch a new program designed to train more ...
May 4, 2022 — RefleXion Medical, a therapeutic oncology company pioneering the use of biology-guided radiotherapy (BgRT) ...
April 26, 2022 — A group of CNIO researchers headed by Manuel Valiente have found a new strategy that could be used in ...


 August 08, 2022
August 08, 2022