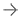November 23, 2021 — The results of a unique two-tiered brain tumor AI challenge were announced today by the Radiological Society of North America (RSNA), Medical Image Computing and Computer Assisted Interventions Society (MICCAI), and the American Society of Neuroradiology (ASNR), who partnered to conduct the challenge. Researchers developed AI models to perform two different tasks using a single dataset of brain magnetic resonance imaging (MRI) scans in competitions staged on separate platforms.
The first challenge task was to build models that perform accurate segmentation and classification of brain tumors in clinically acquired multi-parametric magnetic resonance imaging (mpMRI) scans. This challenge, conducted on the Synapse platform hosted by Sage Bionetworks, is the culmination of a decade of Brain Tumor Segmentation (BraTS) challenges conducted in conjunction with MICCAI, and led by Spyridon Bakas, Ph.D., assistant professor in the Departments of Radiology and Pathology & Laboratory Medicine of the Perelman School of Medicine at the University of Pennsylvania in Philadelphia. This year more than 1,200 participants from five continents registered for this task of the BraTS challenge.
“It is really exciting to see the BraTS challenge, which first occurred in 2012, become the reference benchmark challenge for medical image segmentation with well-curated data of more than 2,000 patients from 37 institutions and more than 1,200 participants,” Bakas said. “We were really honored to have so many expert volunteers believing in this competition and generating gold standard annotations. They create the potential for machine learning algorithms to identify accurately and reproducibly the tumor extent, and hence directly contribute to patient treatment, management and prognosis.”
The second challenge task—introduced this year—involved using the same mpMRI scans to predict the status of a radiogenomic biomarker, a specific genetic sequence observed in analysis of biopsied tumor tissue, known as MGMT promoter methylation, which is an important factor in prognosis and treatment of brain tumors. The Brain Tumor Radiogenomic Classification challenge competition was hosted on the Kaggle platform and drew more than 1,500 participating teams.
“The capability to accurately and non-invasively predict the biochemistry or genetics of a tumor without actually sampling the tissue itself is one of the most compelling potential benefits of medical imaging AI,” said Adam E. Flanders, M.D., neuroradiologist and professor at Thomas Jefferson University Hospital in Philadelphia. “It has the capacity to alter how cancer patients are managed in the future.”
The complete dataset used for these challenge tasks provides a unique resource for AI research in radiology. Built up over a decade of work and substantially enhanced for this year’s challenge, it includes data of 2,040 brain tumor (diffuse glioma) patients from 37 institutions around the world. The data include four distinct structural mpMRI scans for each patient and have been manually annotated with detailed segmentations from more than 60 clinical expert volunteers, predominantly from the ASNR. Contributing sites provided associated MGMT promoter methylation status to facilitate the second task of the challenge. Test set data was retained at the evaluation platform to enable the evaluation of algorithms against unseen hold-out data.
The organizers of the challenge have coordinated with The Cancer Imaging Archive to make publicly available a large majority of the complete dataset early next year. Researchers are invited to continue to submit models for the segmentation task through the Synapse platform, which will provide a sustainable cloud-based platform for open and continuous benchmarking of such algorithms.
The teams who submitted the eight best performing models in each leg of the challenge will share in $60,000 in prize money contributed by the RSNA, Intel Corporation, and Neosoma. The prize-winning teams are:
Segmentation Task
KAIST-MRI-Lab
deepX
mfnv
NVAUTO
FightBrainTumor
Future-Processing-Healthcare
Ngresearch
NPU_PITT
Radiogenomic Classification Task
Tunisia.ai
Minh Phan
Cedric Soares
Leaky Folds
random
Train4Ever
Igor Lashkov
ArturHugo
In addition, an Educational Merit Award was given in each leg of the challenge for the competitors’ clarity in organizing their submissions and explanations of their approach. The winner of the Educational Merit Award for the Segmentation Task is deepX (Yading Yuan). The Educational Merit Award winner for the Radiogenomic Classification Task is Train4Ever (Tung Vu Son, Truong Bui Nhat, Nam Nguyen The, Tuyen Dam Trong, and Khanah Vu Day).
“While challenges encourage development of performant solutions to difficult problems, competitions also provide educational value,” said award review committee chair Katherine P. Andriole, Ph.D., associate professor of radiology at Harvard Medical School, Brigham and Women’s Hospital (BWH) and the Director of Research Strategy and Operations at the MGH (Massachusetts General Hospital) & BWH Center for Clinical Data Science. “For the second year, we acknowledge top-performing entrants for educational merit based on review by a panel of judges of the entrants’ code and supplementary materials including an explanatory video. The quality and educational value of these submissions were excellent. Congratulations to all the competitors!”
The top performers, and all the challenge participants and organizers, will be recognized in an event in the AI Showcase Theater at the RSNA 107th Scientific Assembly and Annual Meeting at McCormick Place in Chicago. The recognition event will be held Nov. 29 from 4:00 to 5:00 p.m. CT.
For more information: RSNA.org

 May 22, 2026
May 22, 2026